Discovery of root anatomy gene may lead to breeding more resilient corn crops
Trait results in roots better able to capture more water and nutrients from soil, need less fertilizer, and withstand drought
A new discovery, reported in a global study that encompassed more than a decade of research, could lead to the breeding of corn crops that can withstand drought and low-nitrogen soil conditions and ultimately ease global food insecurity, according to a Penn State-led team of international researchers.
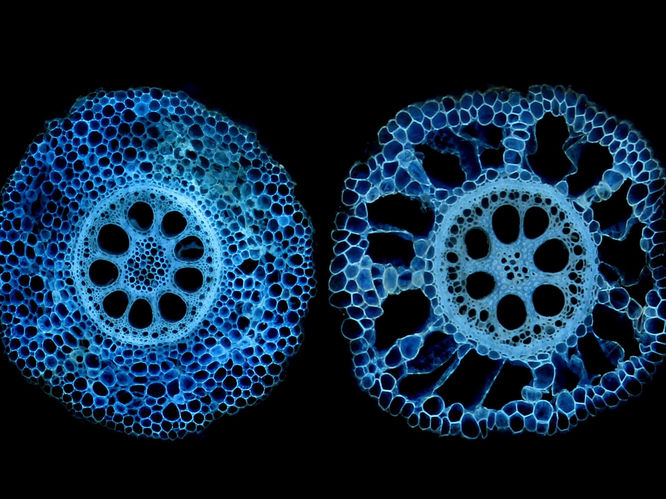
Cross sections of corn roots showing genetic variation for aerenchyma formation, which converts living cells to air space. Now that researchers have found the gene associated with the phenotype, it can be a plant-breeding target.
Tania Galindo Castaneda/Penn State
In findings published March 16 in the Proceedings of the National Academy of Science, the researchers identified a gene encoding a transcription factor – a protein useful for converting DNA into RNA – that triggers a genetic sequence responsible for the development of an important trait enabling corn roots to acquire more water and nutrients.
That observable trait, or phenotype, is called root cortical aerenchyma and results in air passages forming in the roots, according to research team leader Jonathan Lynch, distinguished professor of plant science. His team at Penn State has shown that this phenotype makes roots metabolically cheaper, enabling them to explore the soil better and capture more water and nutrients from dry, infertile soil.
Now, identifying the genetic mechanism behind the trait creates a breeding target, noted Lynch, whose research group in the College of Agricultural Sciences has been studying root traits in corn and beans in the United States, Asia, Latin America, Europe and Africa for more than three decades, with the aim of improving crop performance.
This latest research was spearheaded by Hannah Schneider, formerly a doctoral student and then postdoctoral scholar in the Lynch lab, now assistant professor of crop physiology at Wageningen University & Research, Netherlands. In the study, she used powerful genetic tools developed in previous research at Penn State to accomplish “high-throughput phenotyping” to measure characteristics of thousands of roots in a short time.
Employing technologies such as Laser Ablation Tomography and the Anatomics Pipeline, along with genome-wide association studies, she found the gene — a “bHLH121 transcription factor” — that causes corn to express root cortical aerenchyma. But locating and then validating the genetic underpinnings of the root trait required a prolonged effort, Schneider pointed out.
“We first performed the field experiments that went into this study starting in 2010, growing more than 500 lines of corn at sites in Pennsylvania, Arizona, Wisconsin and South Africa,” she said. “I worked at all those locations. We saw convincing evidence that we had located a gene associated with root cortical aerenchyma.”
But proving the concept took a long time, Schneider related. The researchers created multiple mutant corn lines using genetic manipulation methods such as the CRISPR/Cas9 gene-editing system and gene knockouts to show the causal association between the transcription factor and formation of root cortical aerenchyma.
“It took years not only to create those lines, but also to phenotype them in different conditions to validate the function of this gene,” she said. “We spent 10 years on this, confirming and validating our results, to make sure that this is the gene and the specific transcription factor that controls root cortical aerenchyma formation. Doing this type of work in the field and digging up and phenotyping roots of mature plants was a long process.”
In the paper, the researchers reported that functional studies revealed that the mutant corn line with the bHLH121 gene knocked out and a CRISPR/Cas9 mutant line in which the gene was edited to suppress its function both showed reduced root cortical aerenchyma formation. In contrast, an overexpression line exhibited significantly greater root cortical aerenchyma formation when compared to the wildtype corn line.
Characterization of these lines under suboptimal water and nitrogen availability in multiple soil environments revealed that the bHLH121 gene is required for root cortical aerenchyma formation, according to the researchers. And the overall validation of the bHLH121 gene’s importance in root cortical aerenchyma formation, they propose, provides a new marker for plant breeders to select varieties with improved soil exploration, and thus yield, under suboptimal conditions.
For Lynch, who plans to retire from the Department of Plant Sciences faculty at the end of this year, this research is the culmination of 30 years’ work at Penn State.
“These findings are the result of many people at Penn State and beyond collaborating with us, working over many years,” he said. “We discovered the function of the aerenchyma trait and then the gene associated with it, And, it came about because of technologies that have been devised here at Penn State, such as Shovelomics — digging up roots in the field — Laser Ablation Tomography and Anatomics Pipeline. We put all those together in this work.”
The results are significant, Lynch continued, because finding a gene behind an important trait that's going to help plants have better drought tolerance and better nitrogen and phosphorus capture looms large in the face of climate change.
“Those are super-important qualities — both here in the U.S. and around the world,” he said. “Droughts are the biggest risk to corn growers and are worsening with climate change, and nitrogen is the biggest cost of growing corn, from both a financial and environmental perspective. Breeding corn lines more efficient at scavenging for the nutrient would be a major development.”
Contributing to the research at Penn State were Kathleen Brown, professor of plant stress biology, now retired, Meredith Hanlon, postdoctoral scholar, Department of Plant Science; Stephanie Klein; doctoral student in plant science; and Cody Depew, postdoctoral scholar, Department of Plant Science; and Vai Lor, Shawn Kaeppler and Xia Zhang, Department of Agronomy and Wisconsin Crop Innovation Center, University of Wisconsin; Patompong Saengwilai, Department of Biology, Faculty of Science, Mahidol University, Bangkok, Thailand; Jayne Davis, Rahul Bhosale and Malcolm Bennett, Future Food Beacon and School of Biosciences, University of Nottingham, Loughborough, UK; Aditi Borkar, School of Veterinary Medicine and Science, University of Nottingham, Sutton Bonington, UK.
The U.S. Department of Energy, the Howard G Buffett Foundation, and the U.S. Department of Agriculture’s National Institute of Food and Agriculture supported this research.


























































